Eleutherococcus nodiflorus
Eleutherococcus nodiflorus
1. The products in our compound library are selected from thousands of unique natural products; 2. It has the characteristics of diverse structure, diverse sources and wide coverage of activities; 3. Provide information on the activity of products from major journals, patents and research reports around the world, providing theoretical direction and research basis for further research and screening; 4. Free combination according to the type, source, target and disease of natural product; 5. The compound powder is placed in a covered tube and then discharged into a 10 x 10 cryostat; 6. Transport in ice pack or dry ice pack. Please store it at -20 °C as soon as possible after receiving the product, and use it as soon as possible after opening.

Natural products/compounds from Eleutherococcus nodiflorus
- Cat.No. Product Name CAS Number COA
-
BCN8389
Tritetradecanoin555-45-3
Instructions

-
BCN4600
Kaurenoic acid6730-83-2
Instructions

-
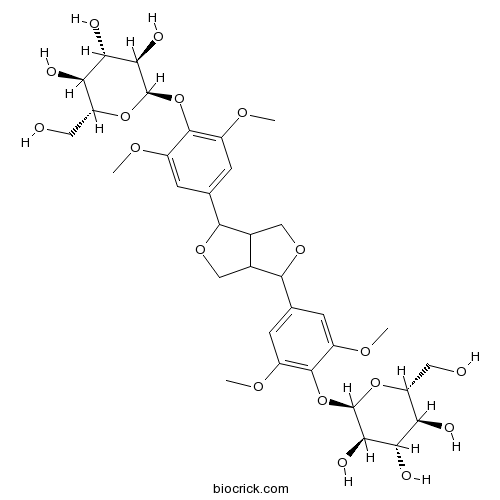
BCN5336
Eleutheroside D79484-75-6
Instructions
-
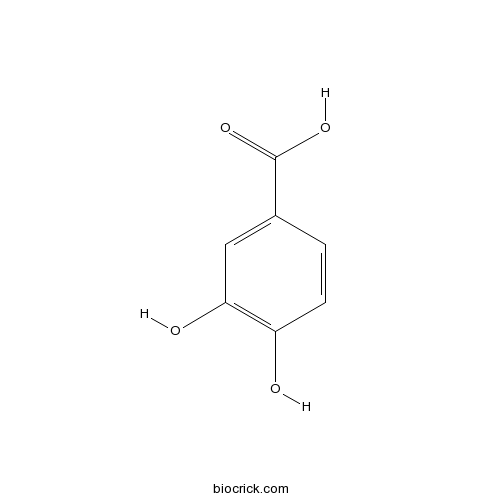
BCN4537
3,4-Dihydroxybenzoic acid99-50-3
Instructions
Internal transcribed spacer 2 barcode: a good tool for identifying Acanthopanacis cortex.[Pubmed: 26500674]
Acanthopanacis cortex has been used in clinical applications for a long time. Considering some historical and geographical factors, Acanthopanacis cortex is easily confused with other herbs in medicine markets, thereby causing potential safety issues. In this study, we used the internal transcribed spacer 2 (ITS2) barcode to identify 69 samples belonging to six species, including Acanthopanacis cortex and its adulterants. The nearest distance, single-nucleotide polymorphisms (SNPs), and neighbor-joining (NJ) tree methods were used to evaluate the identification ability of the ITS2 barcode. According to the kimura-2-parameter model, the intraspecific distance of Eleutherococcus nodiflorus ITS2 sequences ranged from 0 to 0.0132. The minimum interspecific distance between E. nodiflorus and E. giraldii was 0.0221, which was larger than the maximum intraspecific distance of E. nodiflorus. Three stable SNPs in ITS2 can be used to distinguish Acanthopanacis cortex and its closely related species. The NJ tree indicated that the Acanthopanacis cortex samples clustered into one clade, which can be distinguished clearly from the adulterants of this herb. A secondary structure of ITS2 provided another dimensionality to identify species. In conclusion, the ITS2 barcode effectively identifies Acanthopanacis cortex, and DNA barcoding is a convenient tool for medicine market supervision.


