Sinopodophyllum hexandrum
Sinopodophyllum hexandrum
1. The products in our compound library are selected from thousands of unique natural products; 2. It has the characteristics of diverse structure, diverse sources and wide coverage of activities; 3. Provide information on the activity of products from major journals, patents and research reports around the world, providing theoretical direction and research basis for further research and screening; 4. Free combination according to the type, source, target and disease of natural product; 5. The compound powder is placed in a covered tube and then discharged into a 10 x 10 cryostat; 6. Transport in ice pack or dry ice pack. Please store it at -20 °C as soon as possible after receiving the product, and use it as soon as possible after opening.
Natural products/compounds from Sinopodophyllum hexandrum
- Cat.No. Product Name CAS Number COA
-
BCN5608
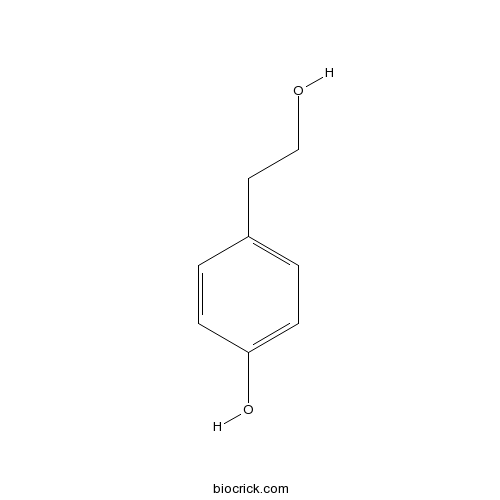
2-(4-Hydroxyphenyl)ethanol501-94-0
Instructions
-
BCN5957
Podophyllotoxin518-28-5
Instructions

-
BCN4537
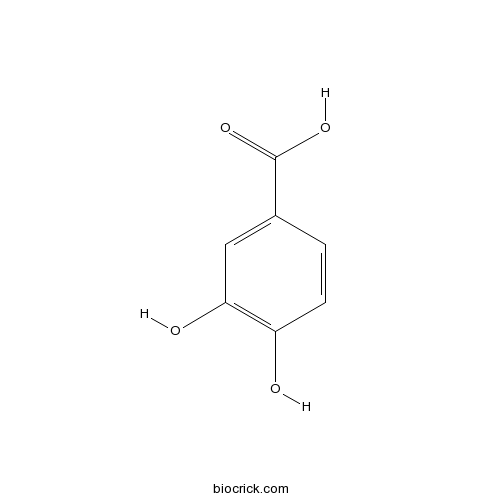
3,4-Dihydroxybenzoic acid99-50-3
Instructions
Plastome organization, genome-based phylogeny and evolution of plastid genes in Podophylloideae (Berberidaceae).[Pubmed: 29981470]
Species of Podophylloideae (Berberidaceae, Ranunculales) are of great pharmacogenetic importance and represent the classic biogeographic disjunction between eastern Asia (EA; 10 ssp.) and eastern North America (ENA; 2 ssp.). However, previous molecular studies of this group suffered from low phylogenetic resolution and/or insufficient marker variability. This study is the first to report whole-plastome sequence data for all 12 species of Podophylloideae (14 individuals) and a close relative, Achlys triphylla. These 15 plastomes proved highly similar in overall size (156,240-157,370 bp), structure, gene order and content, also when compared to other Ranunculales, but also revealed some structural variations caused by the expansion or contraction of the inverted repeats (IRs) into or out of adjacent single-copy regions. Our phylogenomic analysis, based on 63 plastome-derived protein-coding genes (CDS), supported the monophyly of Podophylloideae and its two major genera (EA: Dysosma, EA/ENA: Diphylleia), with Podophyllum peltatum L. (ENA) being more closely related to Diphylleia than to the group's earliest diverging species, Sinopodophyllum hexandrum (EA). Furthermore, within this subfamily/dataset, matK was identified as the fastest evolving gene, which proved to be under positive selection especially in more recently derived, lower-elevation lineages of Dysosma, possibly reflecting an adaptive response to novel environmental (i.e. subtropical compared to higher-elevation/alpine) conditions. Finally, several highly variable noncoding regions were identified in the plastomes of Podophylloideae and Ranunculales. These highly variable loci should be the best choices for future phylogenetic, phylogeographic, and population-level genetic studies. Overall, our results demonstrate the power of plastid phylogenomics to improve phylogenetic resolution, and contribute to a better understanding of plastid gene evolution in Podophylloideae.
[Survival state and fruiting characteristics of endangered medicinal plant Sinopodophyllum hexandrum plant in Tibet plateau].[Pubmed: 29676115]
This study was carried out by the methods of typical plots investigation and laboratory test aiming at analyzing the survival state and fruiting characteristics of three Sinopodophyllum hexandrum populations in different environmental habitats. Meanwhile, it could provide scientific basis for enhancing wild population quantity recovery. The results showed that more population quantity grow in the habitats of large-area gap (Population A) and bushes (Population C) with a majority of the young individuals, while the minor-area gap (Population B) was the opposite. The development tendency of S. hexandrum populations would be the recession in the future. Spatial distribution pattern of populations was clumped at small scales but random at large scales. The indexes of population A and C, as fruit size,the quantity and quality of seeds,germ inability,were all superior to those of population B. Comparing the mainly environment factors of three populations, that favorable environmental factors for vegetative and reproductive growth of S. hexandrum populations were found,especially certain lighting intensity and fertile soil. Therefore, the favorable environmental habitats for S. hexandrum individuals growth could be artificial to promote the recovery and quantities of S. hexandrum populations in the future.
Propioniciclava sinopodophylli sp. nov., isolated from leaves of Sinopodophyllum hexandrum (Royle) Ying.[Pubmed: 28901898]
None
Paenibacillus qinlingensis sp. nov., an indole-3-acetic acid-producing bacterium isolated from roots of Sinopodophyllum hexandrum (Royle) Ying.[Pubmed: 27902261]
A novel indole-3-acetic acid-producing bacterium, designated TEGT-2T, was isolated from the roots of Sinopodophyllum hexandrum collected from the Qinling Mountains in shaanxi province, northwestern China, and was subjected to a taxonomic study by using a polyphasic approach. Cells of strain TEGT-2T were Gram-stain-positive, strictly aerobic, endospore-forming rods and motile by means of peritrichous flagella. Phylogenetic analyses based on 16S rRNA gene sequences showed that strain TEGT-2T was a member of the genus Paenibacillus, exhibiting the highest sequence similarity to Paenibacillus pectinilyticus KCTC 13222T (97.9 %), Paenibacillus frigoriresistens CCTCC AB 2011150T (97.3 %), Paenibacillus ferrarius CCTCC AB 2013369T (96.9 %) and Paenibacillus alginolyticus NBRC 15375T (96.5 %). The only menaquinone detected was MK-7, and the major fatty acid was anteiso-C15 : 0. The major polar lipids were diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine, two unidentified aminophospholipids, two unidentified phospholipids, an unidentified aminolipid and two unidentified lipids. meso-Diaminopimelic acid was detected in the peptidoglycan. The DNA G+C content was 46.6 mol%. DNA-DNA relatedness values for strain TEGT-2T with respect to its closest phylogenetic relatives Paenibacilluspectinilyticus KCTC 13222T and Paenibacillus. frigoriresistens CCTCC AB 2011150T were lower than 40 %. Based on the phenotypic, phylogenetic and genotypic data, strain TEGT-2T is considered to represent a novel species of the genus Paenibacillus, for which the name Paenibacillus qinlingensis sp. nov. is proposed. The type strain is TEGT-2T (=CCTCC AB 2015258T=KCTC 33806T).
Biotechnological interventions for harnessing podophyllotoxin from plant and fungal species: current status, challenges, and opportunities for its commercialization.[Pubmed: 27644897]
Podophyllotoxin is an aryltetralin lignan synthesized in several plant species, which is used in chemotherapies for cancers and tumor treatment. More potent semisynthetic derivatives of podophyllotoxin such as etoposide and teniposide are being developed and evaluated for their efficacy. To meet the ever increasing pharmaceutical needs, species having podophyllotoxin are uprooted extensively leading to the endangered status of selective species mainly Sinopodophyllum hexandrum. This has necessitated bioprospection of podophyllotoxin from different plant species to escalate the strain on this endangered species. The conventional and non-conventional mode of propagation and bioprospection with the integration of biotechnological interventions could contribute to sustainable supply of podophyllotoxin from the available plant resources. This review article is focused on the understanding of different means of propagation, development of genomic information, and its implications for elucidating podophyllotoxin biosynthesis and metabolic engineering of pathways. In addition, various strategies for sustainable production of this valuable metabolite are also discussed, besides a critical evaluation of future challenges and opportunities for the commercialization of podophyllotoxin.
Paenibacillus sinopodophylli sp. nov., a siderophore-producing endophytic bacterium isolated from roots of Sinopodophyllum hexandrum (Royle) Ying.[Pubmed: 27565539]
A Gram-stain-positive, strictly aerobic, rod-shaped, motile and endospore-forming bacterial strain, designated TEGR-3T, was isolated from the roots of Sinopodophyllum hexandrum collected from the Qinling Mountains in Shaanxi Province, China. Strain TEGR-3T produced siderophores and hydrolysed aesculin, starch and CM-cellulose. Phylogenetic analyses based on 16S rRNA gene sequences showed that strain TEGR-3T was a member of the genus Paenibacillus, exhibiting the highest sequence similarity to Paenibacillus endophyticus LMG 27297T (97.3 %) and Paenibacillus castaneae DSM 19417T (97.3 %). MK-7 was the only menaquinone detected and anteiso-C15 : 0 and C16 : 0 were the major fatty acids. The major polar lipids were diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine, two unidentified aminophospholipids, two unidentified phospholipids and an unidentified lipid. The cell-wall peptidoglycan contained meso-diaminopimelic acid as the diagnostic diamino acid. The DNA G+C content was 45.2 mol%. DNA-DNA relatedness values for strain TEGR-3T with respect to its closest phylogenetic relatives Paenibacillus endophyticus LMG 27297T and Paenibacillus castaneae DSM 19417T were lower than 40 %. Based on the phenotypic, phylogenetic and genotypic data, strain TEGR-3T is considered to represent a novel species of the genus Paenibacillus, for which the name Paenibacillus sinopodophylli sp. nov. is proposed. The type strain is TEGR-3T (=CCTCC AB 2016047T=KCTC 33807T).
Genetic diversity and structure of the threatened species Sinopodophyllum hexandrum (Royle) Ying.[Pubmed: 27323174]
Sinopodophyllum hexandrum is an important medicinal plant that has been listed as an endangered species, making the conservation of its genetic diversity a priority. Therefore, the genetic diversity and population structure of S. hexandrum was investigated through inter-simple sequence repeat analysis of eight natural populations. Eleven selected primers generated 141 discernible fragments. The percentage of polymorphic bands was 37.59% at the species level, and 7.66-24.32% at the population level. Genetic diversity of S. hexandrum was low within populations (average HE = 0.0366), but higher at the species level (HE = 0.0963). Clear structure and high genetic differentiation were detected between populations using unweighted pair groups mean arithmetic and principle coordinate analysis. Clustering approaches clustered the eight sampled populations into three major groups, and AMOVA confirmed there to be significant variation between populations (63.27%). Genetic differentiation may have arisen through limited gene flow (Nm = 0.3317) in this species. Isolation by distance among populations was determined by comparing genetic distance versus geographical distance using the Mantel test. The results revealed no correlation between spatial pattern and geographic location. Given the low within-population genetic diversity, high differentiation among populations, and the increasing anthropogenic pressure on this species, in situ conservation measures, in addition to sampling and ex situ preservation, are recommended to preserve S. hexandrum populations and to retain their genetic diversity.
[Community composition and diversity of endophytic fungi from roots of Sinopodophyllum hexandrum in forest of Upper-north mountain of Qinghai province].[Pubmed: 28879736]
High throughput sequencing technology is also called Next Generation Sequencing (NGS), which can sequence hundreds and thousands sequences in different samples at the same time. In the present study, the culture-independent high throughput sequencing technology was applied to sequence the fungi metagenomic DNA of the fungal internal transcribed spacer 1(ITS 1) in the root of Sinopodophyllum hexandrum. Sequencing data suggested that after the quality control, 22 565 reads were remained. Cluster similarity analysis was done based on 97% sequence similarity, which obtained 517 OTUs for the three samples (LD1, LD2 and LD3). All the fungi which identified from all the reads of OTUs based on 0.8 classification thresholds using the software of RDP classifier were classified as 13 classes, 35 orders, 44 family, 55 genera. Among these genera, the genus of Tetracladium was the dominant genera in all samples(35.49%, 68.55% and 12.96%).The Shannon's diversity indices and the Simpson indices of the endophytic fungi in the samples ranged from 1.75-2.92, 0.11-0.32, respectively.This is the first time for applying high through put sequencing technol-ogyto analyze the community composition and diversity of endophytic fungi in the medicinal plant, and the results showed that there were hyper diver sity and high community composition complexity of endophytic fungi in the root of S. hexandrum. It is also proved that the high through put sequencing technology has great advantage for analyzing ecommunity composition and diversity of endophtye in the plant.


